![]()
Bacterial Structure
Bacteria have characteristic shapes. Spherical bacteria, termed cocci, can occur as single organisms or appear in pairs (diplococcus), chains (streptococcus), clusters (staphylococcus), tetrads, or sarcina (cube of eight cocci). Rod-shaped bacteria can be single (bacilli), double (diplobacilli), or in chains (streptobacilli). Some bacteria can have an intermediate shape between a coccus and a bacillus and are therefore termed coccobacilli. The bacteria that cause syphilis (Treponema pallidum) are spiral or corkscrew shaped, and those causing cholera (Vibrio) are comma shaped. Corynebacteria have a club shape, and certain bacilli, including Bacillus anthracis, can contain a spore.
These shapes are maintained by the cell wall of the bacteria composed of peptidoglycan. Bacteria do not have a nucleus surrounded by a membrane, but their chromosome forms a structure usually termed a nucleoid. They do not have membrane-bound organelles; however, they can have storage granules or invaginated plasma membrane inclusions. Their sizes are in the range of micrometers and are about one order of magnitude smaller than eukaryotic cells. However, a bacterium about 600 µm in length and 80 µm in diameter was discovered living in the intestinal tract of surgeonfish in 1993 and was named Epulopiscium fishelsoni.
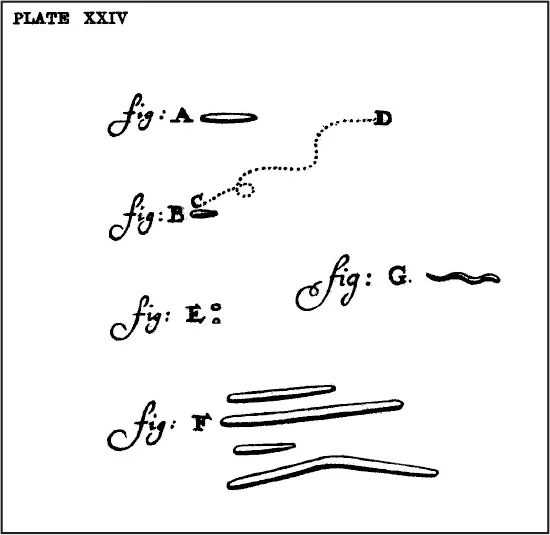
The Dutch scientist Antonie van Leeuwenhoek (1632–1723) is considered to be the first to observe bacteria through his simple microscope. He was inspired by Robert Hooke’s book, Micrographia, to develop a microscope with high enough magnification to examine objects not visible to the naked eye. He utilized a simple biconvex lens that he prepared by flaming soda-lime glass and placed it in between two metal plates. The sample was placed on a pointed metal holder and could be moved into focus by means of screws (Fig 6-1). Van Leeuwenhoek observed red blood cells, the alga Spirogyra from a pond, Giardia from a stool sample, and different bacteria from dental tartar (Fig 6-2). He wrote of his discoveries to the Royal Society in London, and after some initial skepticism, Society members confirmed them in 1673. He was elected as a full member of the Royal Society in 1680.
Fig 6-1 Antonie van Leeuwenhoek’s microscope.
Fig 6-2 Leeuwenhoek’s drawings of the bacteria he observed through his microscope.
Bacteria and other microorganisms can be readily visualized by light microscopy. Because the resolution limit is about 0.2 µm, the optics must be optimized to obtain a clear image. A 100× oil-immersion objective, a 10× eyepiece, and optimal lighting are used to achieve this (Fig 6-3). In bright-field microscopy, it is difficult to visualize small bacteria, so specific dyes are used to stain the bacteria. In dark-field microscopy, a black background is created by blocking the light at the center of the visual field. The light at the periphery is collected by the objective only when it is reflected off the surface of an object, such as a bacterium. Dark-field microscopy is particularly suitable for observing spirochetes. In fluorescence microscopy, the microorganisms are stained with a fluorescent probe, such as a live-dead stain, or a fluorescent antibody. The incident light, which is usually ultraviolet, excites the fluorophore, which then emits light at a higher wavelength (and lower energy). Only the stained microorganisms are seen.
Fig 6-3 (a) Bright-field microscopy. (b) Dark-field microscopy. (c) Fluorescence microscopy. (Reprinted with permission from Ryan and Ray.)
Structurally, bacteria can be classified as being Gram-positive or Gram-negative, based on how they stain with the Gram stain procedure. This procedure was developed serendipitously by the Danish physician Christian Gram in 1884 when an iodine solution spilled over methyl violet-stained lung tissue sections from patients who had died of lobar pneumonia. The currently used procedure involves staining a bacterial sample with crystal violet, setting the dye with Gram’s iodine, decolorizing with alcohol or acetone, and counterstaining with safranin (Fig 6-4). After the decolorization step, the Gram-positive bacteria retain the crystal violet that has been set by Gram’s iodine, whereas the Gram-negative bacteria do not and appear transparent. The counterstain enables the visualization of the latter. A similar procedure is used to stain acid-fast bacteria, including Mycobacterium tuberculosis (Fig 6-5).
Fig 6-4 The Gram staining procedure and the resulting appearance of Gram-positive and Gram-negative bacteria.
Fig 6-5 Comparison of the Gram stain and the acid-fast stain. Four types of bacteria and a neutrophil are shown at each stage of the procedure. In the Gram stain, they are all stained purple by crystal violet and iodine (used as a mordant) (A1). In the acid-fast stain, they are stained red by the carbol fuchsin (B1). After decolorization, Gram-positive bacteria retain the purple stain (A2) and the acid-fast bacteria their red stain (B2). Counterstaining enables the visualization of the other bacteria. In Gram-staining, safranin stains Gram-negative bacteria red (A3). In the acid-fast method, methylene blue stains the remaining organisms blue (B3). (Reprinted with permission from Ryan and Ray.)
Gram-positive bacteria have a single lipid bilayer membrane surrounded by a thick layer of peptidoglycan (also called mucopeptide or murein), which is a mesh of linear polysaccharide co-polymer chains composed of N-acetylglucosamine (GlcNAc) and N-acetylmuramic acid (MurNAc). The chains are cross-linked by peptides (Fig 6-6). The organization of the peptidoglycan layer confers shape to the bacterium. The peptidoglycan also enables the bacterium to withstand osmotic pressure differences across the cytoplasmic membrane of Gram-positive cells and the inner and outer membranes of Gram-negative cells. The β-1,4 glycosidic linkage between GlcNAc and MurNAc acid is the target of the enzyme lysozyme found in tears, saliva, and mucosal surfaces and is directly accessible to the enzyme in Gram-positive bacteria. In Staphylococcus aureus, a Gram-positive coccus, the peptide bridge consists of L-alanine–D-glutamine–L-lysine–D-alanine–D-alanine and is attached to MurNAc (Fig 6-7). The peptides from adjoining polysaccharides are linked by the action of membrane-bound transpeptidases via the pentaglycine bridge attached to the lysine in one peptide and the penultimate D-alanine in the peptide of the adjoining polysaccharide. The terminal D-alanine is removed by a carboxypeptidase. Transpeptidases are penicillin-binding proteins, and they are inhibited by the antibiotic.
Fig 6-6 Schematic representation of the cell wall of Gram-negative (a) and Gram-positive (b) bacteria.
Fig 6-7 The peptidoglycan structure in representative (a) Gram-positive (S aureus) and (b) Gram-negative (E coli) bacteria. (Inspired by Greenwood et al.)
If the components of peptidoglycan are synthesized inside the bacterial cell, how is peptidoglycan assembled outside the cytoplasmic membrane of bacteria?
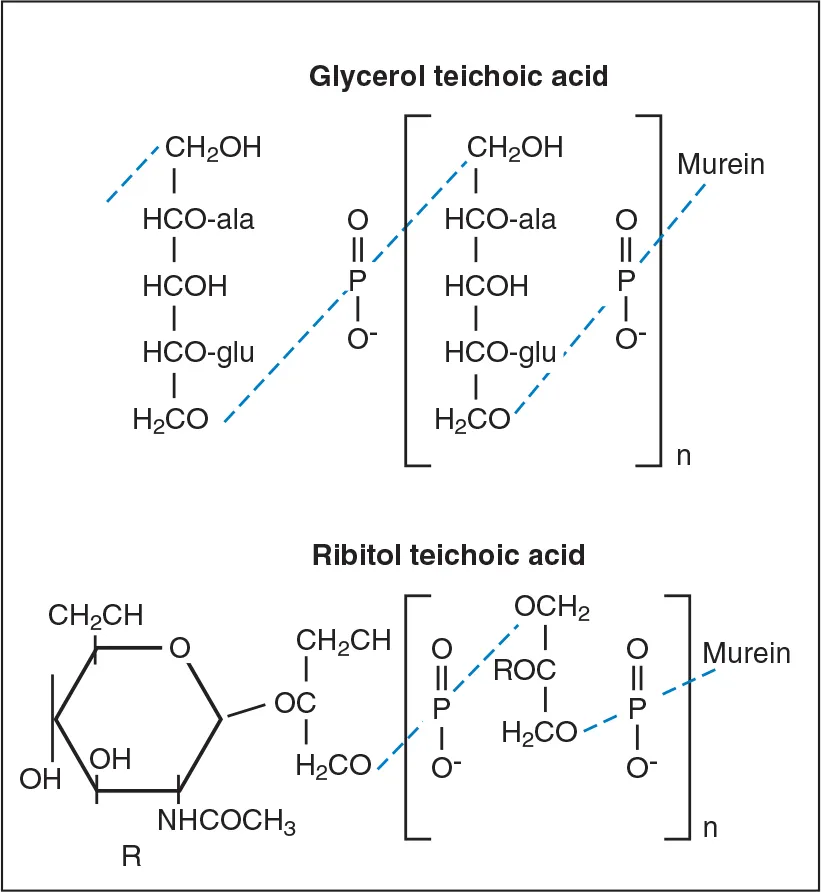
The cell wall of Gram-positive bacteria also contains the polymers, teichoic acids, lipoteichoic acids, and complex (C) polysaccharides. Teichoic acids are polymers of glycerol phosphate or ribitol phosphate (Fig 6-8). Lipoteichoic acids are poly(glycerol phosphates) linked to glycolipid anchors in the cytoplasmic membrane. These molecules facilitate interbacterial attachment and binding to mammalian cell surfaces. They also act as antigens that can be used to distinguish among different serotypes of the microorganism.
Fig 6-8 Glycerol and ribitol teichoic acid. Teichoic acid may be attached covale...