
- 188 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Data Processing Handbook for Complex Biological Data Sources
About this book
Data Processing Handbook for Complex Biological Data provides relevant and to the point content for those who need to understand the different types of biological data and the techniques to process and interpret them. The book includes feedback the editor received from students studying at both undergraduate and graduate levels, and from her peers. In order to succeed in data processing for biological data sources, it is necessary to master the type of data and general methods and tools for modern data processing. For instance, many labs follow the path of interdisciplinary studies and get their data validated by several methods.
Researchers at those labs may not perform all the techniques themselves, but either in collaboration or through outsourcing, they make use of a range of them, because, in the absence of cross validation using different techniques, the chances for acceptance of an article for publication in high profile journals is weakened.
- Explains how to interpret enormous amounts of data generated using several experimental approaches in simple terms, thus relating biology and physics at the atomic level
- Presents sample data files and explains the usage of equations and web servers cited in research articles to extract useful information from their own biological data
- Discusses, in detail, raw data files, data processing strategies, and the web based sources relevant for data processing
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Chapter 1
Mass spectroscopy
Jyotika Rajawat1 and Gagandeep Jhingan2, 1Molecular & Human Genetics Laboratory, Department of Zoology, University of Lucknow, Lucknow, India, 2Valerian Chem Private Limited, New Delhi, India
Abstract
Mass spectroscopy is an analytical technique that identifies biomolecules or proteins present in biological samples and is also useful for studies on protein–protein interactions. The basic principle involves the fragmentation of a compound or molecule into charged species, which are accelerated, deflected, and finally focused on a detector according to their mass and charge ratio. Ion deflection is based on charge, mass, and velocity, ions separation is based on mass to charge (m/z) ratio, and detection is proportional to abundance of ions. This chapter further discusses the anomalies involved in the raw data acquisition, and further processing and interpreting the mass spectrum. For data acquisition, peaks are identified based on preset values from a survey scan and are further taken in for MS/MS analysis. Peptide fragment ion mass is determined for each peak obtained from MS/MS spectrum and subjected to peptide scoring through database search. Protein is identified from the correctly identified peptide scoring.
Keywords
Mass spectrum; orbitrap technology; mass interpretation; protein; phosphoprotein; TPP; PeptideProphet
1.1 Introduction
Mass spectrometry is an analytical technique that identifies biomolecules or proteins present in biological samples and also useful for studying protein–protein interactions. The information obtained can then be further employed to determine protein functions, functional relationships, and the cellular pathways regulated by the proteins. Protein–protein interactions yield insights on functional relationships including, enzyme-substrate, substrates, posttranslational modifying enzymes, protein–coactivator/corepressor, and host–pathogen interactions [1].
Mass spectroscopy–based proteomic workflow consists of the following steps:
- 1. Isolation of protein samples from biological samples and fractionation by 2D gel electrophoresis.
- 2. Fractionated proteins are tryptic digested and resulting peptides are further fragmented.
- 3. Peptides are injected in mass spectrometer (MS) and subjected to quantitative and qualitative analysis.
- 4. The dataset generated is analyzed by appropriate software for identification of the amino acid sequence and for protein quantification.
- 5. Protein identity is assigned based on database search.
1.1.1 Principle
Mass spectroscopy determines the characteristic fragments or ions that arise by organic molecule breakdown. The basic principle involves the bombarding of organic compound with a beam of electrons to generate positively charged ions. Further, the molecular ion is fragmented by utilizing energy of electrons to break the bonds and yield positively charged species or fragment ions. Positive ions or fragment ions formed are further accelerated and deflected in a circular path using magnetic field and then focused on the detector or photographic plate according to their mass and charge. Each ion represents a separate line on plate and recorded in form of peak intensity. Ion deflection is based on charge, mass, and velocity, ions separation is based on mass to charge (m/z) ratio and detection is proportional to abundance of ions.
1.1.2 Instrumentation
The first MS was designed by JJ Thompson in 1912 and was mainly used to study atomic weight of elements and to monitor the abundance of elemental isotopes. Later in the 1950s, with the advent of techniques to vaporize the organic compounds, it became possible to analyze biomolecules with a MS. Following are the main components of a MS as depicted in Fig. 1.1:
- 1. Ionization source
- 2. Analyzer
- 3. Detector
- 4. Data processor/digitizer

The analyzer and detector are kept in a vacuum so that the ions produced do not collide with air molecules and the ion trajectories are maintained at a particular speed. Once the sample is injected in the instrument through the inlet, it gets ionized in the ionization chamber. Ionized species are then extracted into the analyzer, which resolves the ions based upon their mass to charge ratio. Finally these ions are detected by detectors and relative abundance of each resolved ionic species is recorded in the form of mass spectra.
The samples can be injected into the instrument either by direct infusion or through multidimensional chromatographic separation. Analytes can be delivered in staggered manner with the use of chromatographic technique. Gas or liquid chromatography (LC) is commonly used for sample injection in MS. In gas chromatography (GC) the mobile phase used is gas such as hydrogen or helium, which injects the molecules of analytes in the column. The analyte can be effectively separated from the matrix by manipulating the column temperature. GC is limited to compounds that are volatile, heat stable, and lack polar functional groups. GC-MS is often used for comprehensive drug screening. LC is nowadays a preferred method for separation of samples as it is compatible to broad range of analytes and need less for derivatization than GC. The mobile phase uses aqueous and organic solvent in a particular ratio for altering the distribution of analytes and matrix. LC provides fast and effective separation of sample prior to MS analysis [2].
1.1.3 Types of ionization techniques
Different types of ionization methods are employed depending on the type of samples. Volatile samples are subjected to either electron or chemical ionization. Nonvolatile samples are ionized using any of the given techniques:
- 1. Fast atom bombardment
- 2. Matrix assisted laser desorption ionization (MALDI)
- 3. Electrospray
1.1.3.1 Electron ionization
The ionizing agent is high energetic electron emitted from a heated filament and accelerated due to potential difference of around 70 eV. Sample gets ionized by removal of an electron and generation of positively charged ion species.
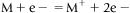
Charged radical cation further gets fragmented into positive ions. Electron ionization (EI) is widely used for volatile organic compounds like oils, hydrocarbons, etc.
1.1.3.2 Chemical ionization
The ionizing agent used is electron but the sample is indirectly ...
Table of contents
- Cover image
- Title page
- Table of Contents
- Copyright
- Dedication
- List of contributors
- Foreword
- Foreword
- Foreword
- Preface
- Acknowledgments
- Chapter 1. Mass spectroscopy
- Chapter 2. Circular dichroism
- Chapter 3. Fluorescence spectroscopy
- Chapter 4. High-throughput sequencing
- Chapter 5. Nuclear magnetic resonance
- Chapter 6. Fourier transform infrared spectroscopy: Data interpretation and applications in structure elucidation and analysis of small molecules and nanostructures
- Chapter 7. Microscopy
- Chapter 8. Principles and applications of flow cytometry
- Chapter 9. Isothermal titration calorimetry
- Chapter 10. Metagenome analysis and interpretation
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Data Processing Handbook for Complex Biological Data Sources by Gauri Misra in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Biotechnology. We have over 1.5 million books available in our catalogue for you to explore.