![]()
1
MicroRNAs: A Brief Introduction
Charles H. Lawrie
Biodonostia Research Institute, San Sebastián, Spain
Nuffield Department of Clinical Laboratory Sciences, University of Oxford, Oxford, UK
I. A Short History of Small RNAs
II. Biogenesis of miRNAs
A. miRNA Nomenclature: What's in a Name?
III. miRNA Function: Controlling mRNA Stability, Degradation, and/or Translation
IV. Regulating the Regulators: miRNA Control and Dysfunction in Disease
A. Genetic Dysregulation of miRNA Expression
B. Epigenetic Regulation
C. Transcription Factors and miRNA Regulatory Networks
D. Regulating miRNA Synthesis and Processing
E. Control of miRNA Function
V. Present and Future Perspectives for miRNAs in Medicine
A. Deciphering the miRNA Targetome: Understanding the Functional Consequences of miRNA Dysregulation in Disease
B. Tip of the Non-Coding RNA Iceberg
C. Are miRNAs Clinically Useful Molecules?
References
Abbreviations
| ADAR | adenosine deaminases that act on RNA |
| Ago | Argonaute |
| ALL | acute lymphoblastic leukemia |
| CLL | chronic lymphocytic leukemia |
| DGCR8 | DiGeorge critical region 8 |
| dsRNA | double-stranded RNA |
| Exp-5 | exportin 5 |
| HITS-CLIP | high-throughput sequencing of RNA isolated by cross-linking immunoprecipitation |
| IP | immunoprecipitation |
| lncRNA | long non-coding RNA |
| miRISC | miRNA RNA interference silencing complex |
| miRNA | microRNA |
| mRNA | messenger RNA |
| ncRNA | non-coding RNA |
| nt | nucleotide |
| PACT | protein activator of the interferon-induced protein kinase |
| PAR-CLIP | photoactivatable-ribonucleoside-enhanced cross-linking and immunoprecipitation |
| PASR | promoter-associated small RNAs |
| piRNA | piwi-interacting RNA |
| PROMPT | promoter upstream transcripts |
| RBP | RNA-binding protein |
| RNAi | RNA interference |
| snoRNA | small nucleolar RNA |
| ssRNA | single-stranded RNA |
| TF | transcription factor |
| tiRNA | tiny RNA |
| TRBP | HIV-1 TAR RNA binding protein |
| tRNA | transfer RNA |
| TSSa-RNA | TSS-associated RNA |
| T-UCR | transcribed ultraconserved regions |
| UTR | untranslated region |
I. A Short History of Small RNAs
The central dogma of molecular biology, first postulated by Francis Crick in 1958 and later refined in 1970, states that biological information flows unidirectionally from DNA to RNA to protein (Crick 1970). This view implies that non-coding RNA (ncRNA) has little or no intrinsic value, despite accounting for more than 90% of eukaryotic transcriptional output (Mattick 2001). Consequently, it is perhaps not surprising that microRNAs (miRNAs) were unknown to the scientific community until very recently. Indeed, it was only in 1993 when the first, what we now know to be a miRNA, was announced by the Ambros and Ruvkun laboratories simultaneously in the December edition of the journal Cell (Lee et al. 1993; Wightman et al. 1993). The Ambros group had identified and cloned a Caenorhabditis elegans developmental regulatory locus, lin-4, that did not contain conventional start and stop codons. Furthermore, introducing mutations that disrupted the putative open reading frame in this 700-nt fragment did not affect function, suggesting that lin-4 did not encode for a protein at all (Lee et al. 1993). At the same time, the Ruvkun lab were working on another temporal regulator of C. elegans, lin-14. They had found that lin-14 was regulated posttranscriptionally via a repeat sequence in the 3′-UTR (untranslated region) of the gene (Wightman et al. 1993). The two labs shared their unpublished findings and realized that the small transcripts of lin-4 (22 nt and 61 nt in length) contained complementary sequences to the 3′-UTR sequence of lin-14, and could regulate this gene via an entirely new regulatory mechanism involving non-coding RNA. However, as lin-4 has no clear homologue outside of worms, the biological significance of this discovery was not realized until many years later.
Although RNA silencing had been known in plants since the beginning of the 1990s (Napoli et al. 1990), the connection with small RNAs was not made until 1999, when the Baulcombe laboratory identified small (25-nt) non-coding RNA species complementary to the target gene that were responsible for gene silencing (Hamilton and Baulcombe 1999). A few months later, it was demonstrated that dsRNA, the trigger for RNA interference (RNAi) (Fire et al. 1998), was sequentially processed into 21–23 nt ssRNA fragments (Zamore et al. 2000). Soon after, another publication from the Ruvkun laboratory described a heterochronic gene of C. elegans, let-7, that controls juvenile to adult transition in larval development (Reinhart et al. 2000). let-7 shared many of the characteristics of lin-4 as it encoded for a small (21 nt) ncRNA transcript that negatively regulated the mRNA of lin family members through complementary RNA-RNA interactions at the 3′-UTR of these genes. Unlike lin-4, however, the sequence of let-7 was found to be conserved in most eukaryotic organisms (Pasquinelli et al. 2000; Lagos-Quintana et al. 2001). Together, these discoveries instigated the start of the miRNA revolution, a term first coined by Lee and Ambros in 2001 (Lee and Ambros 2001). Since this time, over 25,000 miRNAs (including more than 2000 human miRNAs) have been identified from a diverse range of more than 190 different species, including algae, plants, mycetozoa, arthropods, nematodes, protozoa, vertebrates, plants, and viruses (Griffiths-Jones et al. 2006). For a current list of annotated miRNAs, see the miRBase database (http://www.mirbase.org/).
MiRNAs primarily function as posttranscriptional (negative) regulators of gene expression via binding to complementary sequences located mainly within the UTRs of target genes. Because a single miRNA can target several hundred genes, it is believed ∼60% of all human genes are a potential target for miRNA regulation (Friedman et al. 2009). In addition, a single target gene often contains binding sites for multiple miRNAs that can bind cooperatively (Lewis et al. 2003), allowing miRNAs to form complex regulatory control networks. Perhaps, unsurprisingly, miRNAs have been shown to play key regulatory roles in virtually every aspect of biology, including the many physiological and pathological processes described in the chapters of this book.
II. Biogenesis of miRNAs
The majority of human miRNAs are encoded within introns of coding mRNAs, while others are located exgenically, in non-coding mRNAs or within the 3′-UTR sequence of coding mRNA (Rodriguez et al. 2004). MiRNAs are transcribed as 5′-capped large polyadenylated transcripts (pri-microRNA) primarily in a Pol II-dependent manner, although the involvement of Pol-III transcription has also been postulated for miRNAs encoded within Alu repeat sequences (Borchert et al. 2006).
Adenosine deaminases that act on RNA (ADARs) can alter the specificity and binding capacity of miRNA transcripts by changing adenosine bases to inosine post-transcriptionally. For example, ADAR-mediated changes to the pre-miR-151 sequence cause accumulation of the pre-miRNA by blocking Dicer processing (Kawahara et al. 2007a). A selective change to the seed sequence of miR-376 by ADARs causes it to additionally target PRPS1 (Kawahara et al. 2007b). Deep sequencing of mouse brain tissue has identified a number of “edited” miRNAs, increasing the potential repertoire of miRNA targets available for regulation (Chiang et al. 2010).
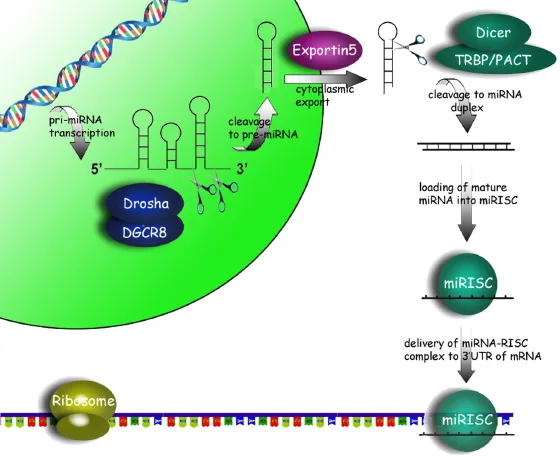
Approximately 40% of human miRNAs are cotranscribed as clusters encoding multiple miRNA sequences in a single pri-microRNA transcript (Altuvia et al. 2005; Hertel et al. 2006). Pri-miRNAs are cleaved within the nucleus by Drosha, an RNaseIII-type nuclease, to form 60–110 nucleotide hairpin structures (pre-microRNA) (Figure 1.1). Drosha by itself possesses little enzymatic activity and requires the cofactor DiGeorge syndrome critical region 8 gene (DGCR8) in humans (Pasha in Drosophila) to form the microprocessor complex (Yeom et al. 2006).
Once produced, pre-miRNAs are exported from the nucleus to the cytoplasm by Exportin-5 (Exp-5) in a Ran-GTP dependent manner (Zeng 2006). The cytoplasmic pre-miRNA is further cleaved by Dicer, another RNaseIII-type enzyme, to form an asymmetric duplex intermediate (miRNA : miRNA*), consisting of the mature miRNA sequence and the antisense miRNA passenger strand (miRNA*). Similar to Drosha, cofactors, such as TRBP and PACT (in humans), are necessary for Dicer activity (Lee et al. 2006). The miRNA : miRNA* duplex is, in turn, loaded into the miRISC complex in which Argonaut (Ago) proteins are the key effector molecules. The strand that becomes the active mature miRNA appears to be dependent upon which has the lowest free energy 5′ end and is retained by the miRISC complex, while the passenger strand is generally believed to be degraded by an unknown nuclease (Khvorova et al. 2003; Schwarz et al. 2003). It should be noted, however, that many miRNA passenger strands are also capable of silencing target transcripts and probably play a more important biological role than was previously realized (Okamura et al. 2008; Ghildiyal et al. 2010).
The loaded miRISC is guided by the mature miRNA sequence (19–24 nucleotide) to complementary sequences located primarily within the 3′-UTR of the target gene mRNA, although binding sites have additionally been identified in both 5′-UTR (Lytle et al. 2007) and coding regions of genes (Tay et al. 2008). In contrast to plant miRNAs that contain extensive regions of complementarity with their target genes, animal miRNAs are only partially complementary and have a propensity to recognize targets via 6–8 nt “seed” sequences, usually located at nt position 2–8 of the 5′-end of the miRNA (Bartel 2009), although sometimes also in the center of the miRNA sequence (Shin et al. 2010). There are rare examples of animal miRNAs (e.g., miR-196 and HOXB8) that do share near-perfect complementarity, resulting in direct cleavage of the ...