
- 384 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Artemia Biology
About this book
Artemia is widely used in both life-sciences research and aquaculture. Although there are over 4000 references regarding Artemia, the literature is widely scattered. Artemia Biology provides a comprehensive, state-of-the-art review of this literature, containing a considerable amount of previously unpublished data. Although all aspects of Artemia biology are covered, the book emphasizes whole-organism approaches. Topics covered include molecular genetics, ontogeny, clonal diversity, mitochondrial DNA-based phylogeny, and comparisons of Artemia and Parartemia (including a taxonomic key to Parartemia species). The book also contains the latest information on Artemia culturing in fertilized ponds and culture tanks, as well as the use of the organism as a food source. Researchers investigating basic biological questions involving molecular genetics, biochemistry, enzymatic and developmental activities, physiology, ecological genetics and adaptation, ecology, and aquaculture production will find this book indispensable.
Tools to learn more effectively

Saving Books

Keyword Search

Annotating Text

Listen to it instead
Information
Chapter 1
Artemia Molecular Genetics
Roberto Marco, Rafael Garesse, Jesús Cruces, and Jaime Renart
TABLE OF CONTENTS
I. Introduction
II. Advantages and Problems of the Artemia System
III. The Genome of Artemia
A. Highly Repeated DNA
B. Intermediately Repeated DNA
1. Ribosomal RNA Genes
2. The 5S RNA-Histone Gene Complex
C. Single Copy DNA
IV. The Genomic Organization of Artemia Mitochondrial DNA
V. DNA Replication, Modification, and Repair
VI. Transcription
VII. Perspectives of Artemia Molecular Genetics
Acknowledgments
References
I. Introduction
Recombinant DNA techniques have made possible the current explosion in the field of molecular genetics.1 The studies of the sixties and early seventies, based primarily on reassociation kinetic analyses, have been superseded by the study of individual genes that can now be isolated, amplified, and manipulated in such ways that a multitude of specific questions can be addressed. The recombinant DNA revolution has also overcome problems such as scarcity of biological materials and the difficulties of in vivo labeling of nucleic acids, etc.
Although the study of Artemia molecular genetics is in early development, several reasons justify its interest. The molecular mechanisms underlying the specific properties and adaptations of Artemia can now be undertaken, such as its capabilities of surviving in extreme environments (e.g., high salt or abnormal pH) or the genetic and molecular aspects of its development. The dissociation between DNA replication and cell division from the onset of cell differentiation and morphogenesis provides an unusual situation in developing organisms. It can be understood as an extreme case of the “midblastula transition”2 that normally takes place while cleavage is going on at full pace. The embryo can be easily induced to resume development and the induction of enzymatic activities or specific mRNAs can be followed with a good reference stage, namely, the encysted early gastrula in which replication, transcription, and translation are completely turned off. (See References 3—5 for previous data on the molecular biology of Artemia.)
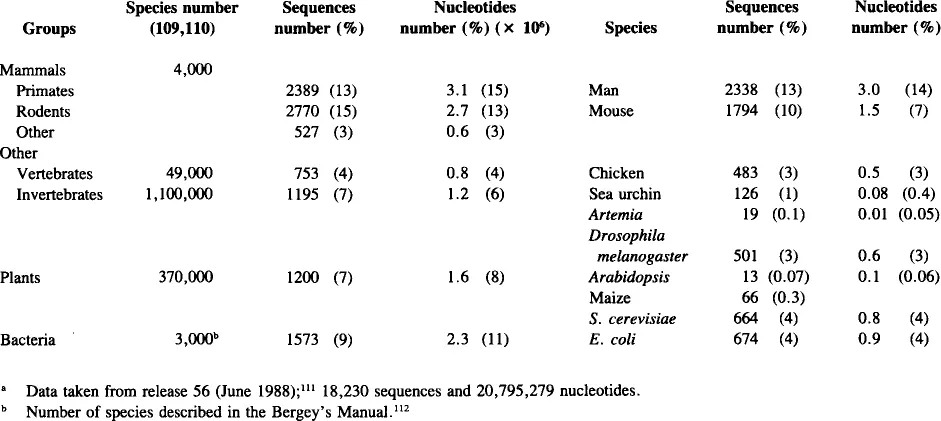
Artemia molecular genetics can fill a gap in evolutionary and comparative biology. Crustacea have not been as extensively studied as other arthropods such as the insect Drosophila melanogaster. In this respect it is interesting to compare the sequence data available from different groups of living organisms (Table 1). The distribution is bimodal with mammals as the best studied group (5686 sequences, 6.4 × 106 nucleotides, 31% of the total) and bacteria (especially E. coli) as the other peak. Among invertebrates, crustaceans represent only 0.02% of the sequences (with 0.01% for Artemia). Insects, and more particularly, Drosophila have been the more extensively investigated invertebrates. The present situation of Artemia is similar to that of Arabidopsis, a plant system that has been presented as an ideal system representative of higher plants.6
Many noninsect invertebrate organisms have been occasionally studied, with nematodes and sea urchins as the best examples. For other classes of arthropods, crustaceans are not well represented, but even less are Chelicerata (0.001% of sequences in the Genebank). The evolutionary distance among these groups is quite large and comparative studies in molecular genetics will help to establish their relationships. In fact, sequence data from a primitive crustacean, such as Artemia, can help to complete the phylogenetic landscape from lower eukaryotes, (e.g., protozoa and fungi), through lower invertebrates (e.g., C. elegans), to the higher invertebrates (e.g., arthropods).
The importance of integrating information obtained from different biological systems using different experimental approaches is increasingly recognized.7 Studying the molecular biology and genetics, the biochemistry and cell biology, and the physiology and development of a particular type of organism in an integrated form and in comparison with other organisms is fundamental in understanding biological diversity. From a molecular point of view, Artemia is the best known crustacean and one of the best known aquatic organisms; thus, it can become the paradigmatic crustacean, a kind of aquatic Drosophila.
The widespread use of Artemia for aquaculture8 further increases the need for basic research on the genus, including genetic manipulation and engineering.
In this review we present the current state of the molecular knowledge of the Artemia genome. We summarize what is known about Artemia nuclear and mitochondrial DNA sequences and the key processes of replication and transcription. This information is discussed in the light of what it is known in other animal organisms, and will serve in a final section to discuss the perspectives about the future of research in this genus.
TABLE 1
Distribution of Sequences in Genbanka
Distribution of Sequences in Genbanka
II. Advantages and Problems of the Artemia System
Artemia presents advantages and disadvantages as an experimental system. The advantages are commercial availability, low cost of the cysts, possibility to synchronize larvae (nauplii) in the laboratory, and a relatively extensive knowledge of the organism’s biochemistry.
The major drawback for the use of Artemia in biochemical and molecular studies is the difficulty in culturing large quantities of particular life cycle stages, including germ cells, early and nondiapausing embryos, nauplii, and adults. Commercial suppliers have also not paid enough attention until recently to the genetic identity of their cysts, which sometimes are not even homogeneous.* Another disadvantage is the relative underdevelopment of classical genetic studies. The high chromosome number and variability of Artemia,9 and the logistical problems involved in keeping separate lines alive complicate the genetic manipulation and use of classical genetic techniques, although cysts can provide a simple and inexpensive way of maintaining stocks. In conclusion, although classical genetic studies are possible, the current level is rather primitive compared to other invertebrate systems, such as Drosophila melanogaster or Caenorhabditis elegans.
Molecular work with Artemia encounters several methodological problems. The impermeability of cysts, the particulate-feeding of nauplii and adults, and the difficulty of rearing large amounts in axenic cultures leads to inefficient in vivo labeling of nucleic acids to specific activities suitable for many experiments. Proteins have been labeled in vivo (see, for instance, References 10—12), but specific activities are low. Furthermore, it is an organism with relatively high amounts of DNA per cell (approximately half that of humans), making it more expensive and tedious than most for screening procedures using molecular techniques. Another problem is the difficulty in obtaining purified nuclei fro...
Table of contents
- Cover
- Title Page
- Copyright Page
- Table of Contents
- Chapter 1 Artemia Molecular Genetics
- Chapter 2 Post-Dormancy Transcription and Translation in the Brine Shrimp
- Chapter 3 Enzyme Activities through Development: A Synthesis of the Activity and Control of the Various Enzymes as the Embryo Matures
- Chapter 4 Normality and Abnormality in Early Development
- Chapter 5 Experimental Biology of Cyst Diapause
- Chapter 6 Morphology of Artemia
- Chapter 7 Ontogeny of Artemia
- Chapter 8 The Extracellular Hemoglobins of Artemia: Structure of the Oxygen Carrier and Respiration Physiology
- Chapter 9 Taxonomy and Population Genetics of Artemia
- Chapter 10 Ecology of Artemia
- Chapter 11 Use of Artemia as Food Source
- Chapter 12 Semi-Intensive Culturing in Fertilized Ponds
- Chapter 13 Production of Artemia in Culture Tanks
- Chapter 14 Anostracans of Australian Salt Lakes, with Particular Reference to a Comparison of Parartemia and Artemia
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.4M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Artemia Biology by Robert A. Browne in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Biology. We have over one million books available in our catalogue for you to explore.