Sivaramakrishnan Rajaraman, Sema Candemir, Zhiyun Xue, Philip Alderson, George Thoma, and Sameer Antani
1.1 Introduction
Tuberculosis (TB) is an infectious disease caused by a rod-shaped bacterium called Mycobacterium tuberculosis. According to the 2018 World Health Organization (WHO) report, there were an estimated 10 million new TB cases, but only 6.4 million (64%) were reported for treatment [1]. Countries including India, China, Pakistan, South Africa, and Nigeria accounted for more than 60% of the people suffering from the infection. A chest X-ray (CXR), also called chest film or chest radiograph, is the most common imaging modality used to diagnose conditions affecting the chest and its contents [2, 3, ]. CXR diagnosis has revolutionized the field of TB diagnostics and is extremely useful in establishing a plausible diagnosis of the infection. Clinicians initiate treatment for the infection based on their judgment of these radiology reports. Posterior-anterior (PA) and lateral CXR projections are routinely examined to diagnose the conditions and provide diagnostic evidence [4]. Figure 1.1 (a)–(e) shows some instances of abnormal and normal CXRs.
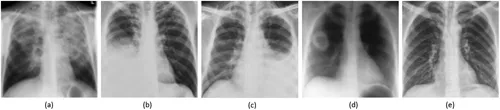
FIGURE 1.1
CXRs: (a) hyper-lucent cystic lesions in the upper lobes, (b) right pleural effusion, (c) left pleural effusion, (d) cavitary lung lesion in the right lung, and (e) normal lung.
With significant advancements in digital imaging technology there is an increase in the use of CXRs for TB screening. However, there is a lack of expertise in interpreting radiology images, especially in TB endemic regions, which adversely impacts screening efficacy [5], an ever-growing backlog and increased opportunity for disease spread. Also, studies show that there is a high degree of variability in the intra-reader and inter-reader agreement during the process of scoring CXRs [6]. Thus, current research is focused on developing cost-effective, computer-aided diagnosis (CADx) systems that can assist radiologists in interpreting CXRs and improve the quality of diagnostic imaging [7]. These systems are highly competent in reducing intra-reader/inter-reader variability and detection errors [8–11]. There are several prior approaches using traditional image analysis and machine learning (e.g. support vector machine [SVM]) that are valuable for providing background on CADx tools for CXR analysis [12–15]. The reader is referred to these as background. They are promoted as a convenient tool to be used in systematic screening and triaging algorithms due to the increased availability of digital radiography, which presents numerous benefits over conventional radiography, including enhanced image quality, safety, and reduced operating expenses [16]. CADx tools have gained immense significance; the appropriate use and advancing of these systems could improve detection accuracy and alleviate the human burden in screening. Earlier CADx studies were based on image segmentation and textural feature extraction with grey-level co-occurrence matrix [17]. A CADx system for TB detection was proposed by Van Ginneken et al. [18], who used multi-scale feature banks for feature extraction and a weighted nearest-neighbor classifier for classification of TB-positive and normal cases. The study demonstrated area under a curve (AUC) values of 0.986 and 0.82 on two private CXR datasets. A technique based on pixel-level textural abnormality detection was proposed by Hogeweg et al. [19] to obtain AUC values between 0.67 and 0.86. However, a comparative study of the proposed methods was hampered due to unavailability of public CXR datasets. Jaeger et al. [20] made available the public CXR datasets for TB detection, followed by Chauhan et al. [21], who helped to evaluate the proposed techniques on public datasets. Melendez et al. proposed the multiple instance learning methods for TB detection, which used moments of pixel intensities as features to be classified by an SVM classifier [22]. The authors obtained AUC between 0.86 and 0.91 by evaluating on three private CXR datasets. Jaeger et al. [5] proposed a combination of standard computer vision algorithms for extracting features from chest radiographs. The study segmented the region of interest (ROI) constituting the lungs, and extracted the features using a combination of algorithms that included a histogram of oriented gradients (HOG), local binary patterns (LBP), Tamura feature descriptors, and other algorithms. A binary classifier was trained on these extracted features to classify normal and TB-positive cases. CADx software based on machine learning (ML) approaches using a combination of textural and morphological features is also commercially available. This includes CAD4TB, a CADx software from the Image Analysis Group, Nijmegen, Netherlands that reported AUC ranging from 0.71 to 0.84 in a sequence of studies performed in detecting pulmonary abnormalities [23]. Another study achieved AUC of 0.87 to 0.90 by using an SVM classifier to classify pulmonary TB from the normal instances using texture and shape features [24]. However, the performance of textural features was found to be inconsistent across the imaging modalities. These features performed well as long as they were able to correlate with the disease, but delivered sub-optimal performance in instances when there was an overlapping of anatomical sites and images having complex appearances [25]. Feature descriptors such as bag-of-words (BOW) were also used in discriminating normal from pathological chest radiographs [26]. The method involves representing an image using a bag of visual words, constructed from a vocabulary of features extracted by local/global feature descriptors. A majority of CADx studies used handcrafted features that demand expertise in analyzing the images and account for variability in the morphology and texture of the ROI. On the other hand, deep learning (DL) models learn hierarchical layer-wise representation to model data at more and more abstract representations. These models are also known as hierarchical ML models that use a cascade of layers of non-linear processing units for end-to-end feature extraction and classification [27]. Convolutional neural networks (CNN), a class of DL models, have gained immense research prominence in tasks related to image classification, detection, and localization, as they deliver promising results without the need for manual feature selection [28]. Unlike kernel-based algorithms such as SVMs, DL models exhibit improved performance with an increasing number of training samples and computational resources [29].
Medical images contain visual representations of the interior of the body that aid in clinical analysis and medical intervention [30]. These images are specific to the internal structures of the body and have less in common with natural images. Under these circumstances, a customized CNN, specifically trained on the underlying biomedical imagery, could learn “task-specific” features to aid in improved accuracy. The parameters of a custom model could be optimized for improvement in performance. The learned features and salient network activations could be visualized to understand the strategy the model adapts to learn these task-specific features [31]. However, the performance improvement of customized CNNs comes at the cost of huge amounts of labeled data, which are difficult to obtain, particularly in biomedical applications. Transfer Learning (TL) methods are commonly used to relieve issues with data inadequacy where DL models are pre-trained on large-scale datasets [32]. These pre-trained models could be used either as an initialization for visual recognition tasks or as feature extractors from the underlying data [33]. There are several pre-trained CNNs available, including AlexNet [34], VGGNet [35], GoogLeNet [36], ResNet [37], etc., which transfer knowledge gained from learning a comprehensive feature set from the large-scale datasets to the underlying task and serve as feature extractors in an extensive range of visual recognition applications, outperforming the handcrafted features [38]. Study of the literature reveals the use of pre-trained CNNs...