
- 338 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Ecophysiology of Coniferous Forests
About this book
Conifers--pine, fir, and spruce trees--are dominant species in forests around the world. This book focuses on the physiology of conifers and how these physiological systems operate. Special consideration is devoted to the means by which ecophysiological processes influence organismal function and distribution. Chapters focus on the genetics of conifers, their geographic distribution and the factors that influence this distribution, the impact of insect herbivory on ecophysiological parameters, the effects of air pollution, and the potential impact that global climatic changes will have upon conifers. Because of the growing realization that forests have a crucial role to play in global environmental health, this book will appeal to a developing union of ecologists, physiologists and more theoretically minded foresters.
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
1
Genetics and the Physiological Ecology of Conifers
Jeffry B. Mitton
Publisher Summary
This chapter focuses on the genetic variability associated with the physiological differences among genotypes in Rocky Mountain conifers. It illustrates genetically based differences in physiology by considering variation among genotypes in survival, growth, and resistance to herbivores and presents the mechanistic studies needed to understand the relationships between genetic and physiological variation. In addition to the marked elevation distribution of conifer population size in mountainous regions—especially in the Rocky Mountains—many conifers have other life-history traits and ecological circumstances that promote high levels of genetic variation. Data collected from numerous provenance studies and allozyme surveys confirm this phenomenon. Provenance surveys show that Douglas fir, ponderosa pine, and lodgepole pine populations in the Rocky Mountains differ by 300 m in elevation and likely exhibit adaptation to different environments, while allozyme studies in conifers reveal populations with high levels of variation within populations and only slight variation among populations. Molecular techniques being employed these days help examine variation in the nuclear, mitochondrial, and chloroplast genomes. Provenance studies reveal adaptation to local environments in most of the Rocky Mountain conifers but don’t reveal the localization of the genes that produce morphological, physiological, and phenological responses to environmental gradients in conifer populations. Examination of the physiological and demographic consequences of allozyme loci and variation of nuclear, mitochondrial, and chloroplast DNA help answer whether mitochondrial and chloroplast genomes contribute to the adaptive variations in conifers.
Natural selection acts on the diversity of genotypes, adapting populations to their specific environments and driving evolution in response to changes in climate. Genetically based differences in physiology and demography adapt species to alternate environments and produce, along with historical accidents, the present distribution of species. The sorting of conifer species by elevation is so marked that conifers help to define plant communities arranged in elevational bands in the Rocky Mountains (Marr, 1961). For these reasons, a genetic perspective is necessary to appreciate the evolution of ecophysiological patterns in the coniferous forests of the Rocky Mountains.
The fascinating natural history and the economic importance of western conifers have stimulated numerous studies of their ecology, ecological genetics, and geographic variation. These studies yield some generalizations, and present some puzzling contradictions. This chapter focuses on the genetic variability associated with the physiological differences among genotypes in Rocky Mountain conifers. Variation among genotypes in survival, growth, and resistance to herbivores is used to illustrate genetically based differences in physiology, and to suggest the mechanistic studies needed to understand the relationships between genetic and physiological variation.
I Conifers Have High Levels of Genetic Variation
Evolutionary theory predicts that species that have large population sizes and broad geographic ranges will have high levels of genetic variation. This prediction is based on several relationships. First of all, the number of new mutations arising each generation increases with population size. In addition, the rate of genetic drift is inversely proportional to population size, so that more mutations are expected to accumulate in large populations. Jointly, these relationships predict that genetic variation will increase with population size. For example, the expected number of alleles at a locus k is k = 4Nu + 1, where N is the population size and u is the neutral mutation rate (Hard and Clark, 1989). Similarly, the predicted heterozygosity, H, also increases with population size:
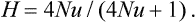
In addition to population size, many conifers have other life-history traits and ecological circumstances that promote high levels of genetic variation. For example, many species of conifers, such as ponderosa pine (Pinus ponderosa), lodgepole pine (Pinus contorta), and Douglas fir (Pseudotsuga menziesii), have immense geographical ranges that place them in a wide variety of natural environments. The degree of environmental variation experienced by a species is expected to increase with the geographic range of the species, and the selection among heterogeneous environments is expected to enhance genetic variation (Hedrick et al., 1976; Hedrick, 1986; Gillespie and Turelli, 1989). We can easily imagine that the Douglas fir growing in sympatry with pinyon pine (Pinus edulis) in the Chiricahua Mountains of southern Arizona will be selected to have traits very different from those favored in the Douglas fir growing in sympatry with Sitka spruce (Picea sitchensis) in the temperate rain forest of the Olympic Peninsula.
Forest biologists detect genetic variation with diverse methods. Trees planted as seeds into common gardens in provenance studies yield estimates of heritability for virtually any character that can be measured. Heritability is the proportion of total phenotypic variation attributable to genetic variation. By planting seeds from diverse sources into a common garden, the environmental differences among collection sites are eliminated, and therefore apparent differences can be attributed to genetic differences among collection localities. The use of several common gardens in contrasting environments allows inferences of how genotypes respond to a diversity of environmental conditions. Although common garden studies reveal differences among genotypes and patterns of performance along environmental gradients, individual genes are not identified. Specific genes or sequences of DNA from seeds and needles collected in the field can be examined with electrophoretic surveys of proteins and with a variety of molecular techniques. Surveys of electrophoretically detectable genetic variation of proteins, or allozyme variation, have been used to measure the genetic variation and describe the geographic variation of many species of conifers (Hamrick et al., 1979; Hamrick and Godt, 1990; Loveless and Hamrick, 1984; Mitton, 1983). The DNA from the nucleus, from mitochondria, and from chloroplasts can be sequenced, or examined for restriction fragment length polymorphisms (RFLPs) (Wagner, 1992a,b), including examination of the variable number of tandem repeats (VNTRs), also popularly referred to as DNA fingerprints.
A Provenance Studies
Provenance studies of many species of conifers have identified substantial proportions of genetic variation for characters such as germinability, growth, disease resistance, times of bud break and bud set, susceptibility to frost damage, and growth form (Wright, 1976). One indication of the magnitude of variation among individuals is the range of values for maximum photosynthetic capacity, gleaned from the review of Ceulemans and Saugier (1991) (Table I). Clear differences distinguish species, but these data illustrate that there is substantial variation within species as well. In both sweet chestnut and black cottonwood, the maximum photosynthetic capacity varies by a factor of two among individuals.
Table I
Range of Average Photosynthetic Capacities for a Variety of Treesa
| Species | Common name | Range (μmol/sec/m2) |
| Castanea sativa | Sweet chestnut | 8–16 |
| Fraxinus pennsylvanica | Green ash | 20–25 |
| Populus tremuloides | Quaking aspen | 20–22 |
| Populus trichocarpa | Black cottonwood | 14–30 |
| Populus euramericana | Euramer... |
Table of contents
- Cover image
- Title page
- Table of Contents
- Physiological Ecology: A Series of Monographs, Texts, and Treatises
- Copyright
- Contributors
- Preface
- Chapter 1: Genetics and the Physiological Ecology of Conifers
- Chapter 2: Long-Term Records of Growth and Distribution of Conifers: Integration of Paleoecology and Physiological Ecology
- Chapter 3: Plant Hormones and Ecophysiology of Conifers
- Chapter 4: Ecophysiological Controls of Conifer Distributions
- Chapter 5: Physiological Processes during Winter Dormancy and Their Ecological Significance
- Chapter 6: Ecophysiology and Insect Herbivory
- Chapter 7: Leaf Area Dynamics of Conifer Forests
- Chapter 8: Causes and Consequences of Variation in Conifer Leaf Life-Span
- Chapter 9: Response Mechanisms of Conifers to Air Pollutants
- Chapter 10: Potential Effects of Global Climate Change
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Ecophysiology of Coniferous Forests by William K. Smith, Jacques Roy in PDF and/or ePUB format, as well as other popular books in Technology & Engineering & Botany. We have over 1.5 million books available in our catalogue for you to explore.