Genome sequencing enables scientists to study genes over time and to test the genetic variability of any form of life, from bacteria to mammals. Thanks to advances in molecular genetics, scientists can now determine an animal's degree of inbreeding or compare genetic variation of a captive species to wild or natural populations. Mapping an organism's genetic makeup recasts such terms as biodiversity and species and enables the conservation of rare or threatened species, populations, and genes.
By introducing a new paradigm for studying and preserving life at a variety of levels, genomics offers solutions to previously intractable problems in understanding the biology of complex organisms and creates new tools for preserving the patterns and processes of life on this planet. Featuring a number of high-profile researchers, this volume introduces the use of molecular genetics in conservation biology and provides a historical perspective on the opportunities and challenges presented by new technologies. It discusses zoo-, museum-, and herbarium-based biological collections, which have expanded over the past decade, and covers the promises and problems of genomic and reproductive technology. The collection concludes with the philosophical and legal issues of conservation genetics and their potential effects on public policy.

eBook - ePub
Conservation Genetics in the Age of Genomics
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Conservation Genetics in the Age of Genomics
About this book
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
Subtopic
EcologyIndex
Biological Sciences | Part I |
Perpectives on the Union of Conservation and Genetics
Conservation genetics has seen an expansion of goals and objectives over the past decade. This came about mostly as a result of the infusion of genomic technology into how we examine the genetics of natural and captive populations. Connecting the original goals of conservation genetics with the expanded potential of genomics is an important process, and the three chapters in this section attempt to make this connection. As an introductory chapter, we reprint from Nature Review Genetics a review by two of the organizers of the New York meeting (Rob DeSalle and George Amato). This chapter describes the incorporation of molecular information into conservation biology and discusses how such approaches have contributed to the expansion of conservation genetics. In chapter 2 William Conway provides an impassioned perspective on the real problems in conservation biology that can be addressed using genetic techniques. Most important, Conway suggests that conservation genetics must focus on area and ecosystem issues; focusing on broken ecosystems and marginalized species can be the best approach to this larger conservation biology goal. In the third chapter, George Amato examines the current state of conservation genetics and biology and concludes that a new paradigm for connecting ecological, life history, and area-based issues with genetics is essential for the field to remain vibrant and useful to conservation.
 | Rob DeSalle and George Amato Adapted with permission from DeSalle, R. and G. Amato. 2004 . The expansion of conservation genetics. Nature Reviews. Genetics 5:702–12. |
The Expansion of Conservation Genetics
Conservation biology has been accurately described as a crisis discipline. Much like the human disease crisis disciplines of HIV biology and cancer biology, conservation biology requires an immediate understanding of the patterns and processes that make it such a critical subject. The urgency of the crises that are the subject of conservation biology are manifest in the large number of species facing imminent extinction. In fact, the last two centuries of human activity have been described as one of the most severe periods of mass extinction of all time, challenging even the most extreme periods of extinction in the long past as a period in which the majority of living species on Earth perish (Jablonski, 1986; Lande, 1988; Soulé, 1986). In addition, because conservation biologists have to make rapid decisions based on currently available data, there is a heightened sense of crisis in the discipline.
Crisis disciplines often see periods of expansion of the toolbox used to address the dilemmas posed by the forces causing the crises. Many of these tools are added to help researchers cope with the immediacy of the problems they are facing and allow them to rapidly and efficiently collect data on problems needing quick and even immediate attention. In conservation biology these tools have advanced accordingly. Some examples of this toolbox expansion include the use of global positioning systems (GPSs), satellite-based imaging of areas, the inclusion of mathematical advances in conservation theory, and perhaps the most visible: genetics. One of the factors involved in the inclusion of genetics in conservation biology over the past decade has been the proliferation of technology in genomics, systematics, and population biology, and conservation geneticists have scrambled to keep up with the pace of progress.
This essay attempts to articulate the current structure and scope of conservation genetics and to demonstrate the utility of this structure in modern conservation biology. The efficiency of this structure in conservation decision making is increased by the expanded genomic technologies in data acquisition, storage, and analysis. The improved precision and quantity of data on endangered species that can now be used in conservation genetics allows us to examine issues of captive species breeding, species boundary problems, and conservation forensics, three topics that will be the focus of this review. Another emerging area of conservation genetics concerns studies that combine genetic methods with ecological and landscape approaches. Such studies can provide conservation biologists with a much more accurate picture of the complex systems they work on.
Conservation biology, and conservation genetics with it, is expanding to include many subdisciplines and approaches, including genomics and high-throughput methods of data acquisition, and this expansion is better enabling the field to address the urgent problems involved in managing endangered species and critical areas. Significant challenges remain, and these include understanding that the context dependency of conservation decisions can be clarified by genetic information. Genetics can provide conservationists with unprecedented precision and elucidate the genetic parameters on which many decisions are made. Most importantly, conservation genetics must be placed in the context of the difficulties of working across political boundaries, amid economic challenges, and in the face of the complexity of using science to inform management decisions.
The Scope of Conservation Genetics
The expansion of the scope of conservation genetics over the past decade has been brought about by the infusion of high-throughput DNA sequencing and genotyping technology. Conservation geneticists have followed the trends of technique usage outlined by Schlotterer (2004) for population genetics as a whole. These techniques are well suited to enhancing our understanding of the evolutionary processes of endangered populations and species. An examination of the intensity of the use of various techniques applicable at the population level (Schlotterer, 2004) in animals indicates that although some techniques, such as random amplification of polymorphic DNA (RAPD), minisatellites, restriction fragment length polymorphism (RFLP), and allozymes, are either extinct or sparingly used in population-level studies, four methods—amplified fragment length polymorphism (AFLP), DNA sequencing, single nucleotide polymorphism (SNP) analysis, and microsatellites—make up most of the techniques used at this level for animals, and plant studies have continued to use AFLP, RAPD, and another DNA variation technique called inter–simple sequence repeat (ISSR; Godwin et al., 1997). The discovery of pattern in conservation genetics has also followed the development of techniques in systematics incorporating high-throughput sequencing and SNP analysis. On the whole, then, conservation geneticists have closely followed the development of genetic and genomic technology and have voraciously incorporated SNPs (Aitken et al., 2004; Brumfield et al., 2003; Zhang and Hewitt, 2003) and microsatellite variation as the major tools of their trade and are looking for more rapid and efficient screening techniques for genetic variability. Finally, some conservation geneticists have contemplated the utility of microarrays and quantitative trait mapping in conservation biology (Gibson, 2002; Purugganan and Gibson, 2003).
The scope of genetics in conservation biology is diverse and has been addressed in several publications (see table 1.1; Avise, 2004; Avise and Hamrick, 1996; Frankham et al., 2003; Spellerberg, 1997; Young et al., 2000). Rather than attempt a comprehensive review of these usages, we instead attempt to place the major approaches to understanding pattern and process in a conservation context. To do this we first examine the early challenges to the infusion of genetics into conservation decision making and then discuss the present and future of pattern and process discovery in conservation genetics.
Table 1.1 The roles of conservation genetics are listed in the first column. The middle column indicates whether the listed role uses pattern or process from the genetic inference. The last column indicates the subdiscipline of evolutionary biology—systematics or population genetics—that is the major source of techniques to accomplish the role.
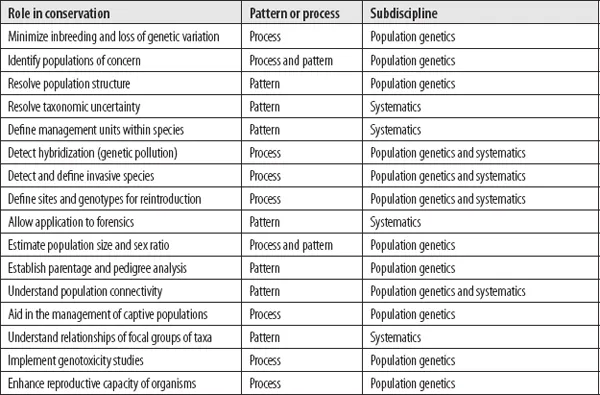
Source: The list of roles is partially after Allendorf and Leary (1986).
Early Challenges
The expansion of the toolbox of conservation biology in the late 1980s to include molecular evolutionary genetic techniques and markers was immediately met with three challenges to the relative merit and priority of genetics in the discipline. The first was a cogent direct challenge to the relevance of genetics to demographic issues in conservation biology by Lande (1988). In a landmark paper, Lande pointed out that demographic factors (the biology of population growth and life history) were much more important in explaining extinction than any of the genetic factors that could be incorporated into a theory of conservation biology. These demographic factors were viewed as swamping out most of the genetic effects ascribed to extinction. Lande’s challenge to conservation genetics was a healthy one: It opened the way for a better-defined role of the concepts of inbreeding and genetic variation in the discipline. It is also clear that population genetics without demographics in conservation usually leads to less than useful recommendations (Ballou and Lacy, 1995). However, careful integration of demographic and fine-grained genetic approaches can often allow strong conservation inferences.
The second challenge came on the pattern side of the conservation genetics toolbox. The idea of conservation units was challenged by Ryder (1986), and the term evolutionarily significant unit (ESU) was imbedded in the conservation genetics literature as a result of healthy debate after Ryder’s original challenge to subspecies definition and, more importantly, to the utility of subspecies definitions in conservation biology (Amato, 1991; O’Brien and Mayr, 1991; Waples, 1991). This problem of unit designation in conservation is often set at the interface of population genetics and systematics, because the goal is to discover species units. These debates, at both levels of resolution, resulted in a more applied focus in conservation genetics research. Clearly articulating the goals of a specific conservation problem provided better guidance to the selection of techniques, tools, and theory.
The third challenge was a posthumously published paper by Caughley (1994) that suggested that too much focus on technical approaches to conservation (including conservation genetics) had resulted in neglect of more important issues such as habitat threat and disease. This criticism was answered by the enormous expansion in the use of conservation genetics in ecology and applied wildlife management (Hedrick et al., 1996) and the realization that genetics could aid in landscape ecology approaches.
Pattern and Process: The Targets of Conservation Genetics
The most significant result of debate on these three challenges was to define the roles of conservation genetics in elucidating genetic and evolutionary processes and delineating the patterns relevant to managing endangered populations. Understanding that genetics leads us to a better picture of pattern and process in endangered species defines the most important roles of conservation genetics. The first is to aid in a more precise description and understanding of the processes that gave rise to the current state of an endangered population or species. This role is most important because the identification of factors such as inbreeding depression, breeding effective population size, minimum viable population size, and levels of genetic variation and gene flow in natural populations (Allendorf and Leary, 1986; Ralls et al., 1988; Templeton and Read, 1984; Waples, 1989) lends greater definition to the processes that affect endangered populations and also indicates immediate genetically based responses to the detrimental effects of these processes.
The tools used to analyze intrapopulation and genealogical problems have been summarized in several publications (Glaubitz et al., 2003; Goodnight and Quellar, 1999; Petit et al., 2002; Ritland, 2000; Vekemens and Hardy, 2004). Ex situ population genealogical and inbreeding analysis is a particularly important area where these methods are applied, and several studies making recommendations concerning breeding programs have arisen from these kinds of pedigree analysis studies (Geyer et al., 1993; Lacy, 2000a; Miller et al., 2003; Russello and Amato, 2001, 2004). The urgency of making the best possible genetic decisions in breeding programs of endangered species is exemplified by several studies in which pedigrees inform matchmaking. In ...
Table of contents
- Cover
- Half title
- Series Page
- Title
- Copyright
- Contents
- List of Illustrations
- List of Tables
- Foreword: Sydney Brenner, The Continuity of Genomes and Genetic Resources for the New Century
- Acknowledgments
- General Introduction
- Part I. Perspectives on the Union of Conservation and Genetics
- Part II. Conservation Genetics in Action: Assessing the Level and Quality of Genetic Resources in Endangered Species
- Part III. Saving Genetic Resources
- Part IV. Genomic TechPartlogy Meets Conservation Biology
- Part V. Policy, Law, and Philosophy of Conservation Biology in the Age of Genomics
- Further Reading
- List of Contributors
- Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Conservation Genetics in the Age of Genomics by George Amato,Rob DeSalle,Oliver A. Ryder,Howard C. Rosenbaum, George Amato, Rob DeSalle, Oliver Ryder, Howard Rosenbaum in PDF and/or ePUB format, as well as other popular books in Biological Sciences & Ecology. We have over 1.5 million books available in our catalogue for you to explore.