1.1 Statement of the Problem of Structure Elucidation
The first reports devoted to computer-assisted structure elucidation (CASE) were published by four independent groups of researchers1–4 in the late 1960s. Prior to describing CASE methods, we will consider the complex nature of the structure elucidation problem.
It is necessary to distinguish two different analytical problems that are associated with the structure elucidation of molecules. The first relates to the identification of a molecule that is assumed to be known already and whose physicochemical characteristics, specifically the associated spectral data, are included into collections of reference data. In this case the solution is found by searching through the reference data and comparing the data measured for the unknown with the available reference data. Most frequently the search is performed using one or more of the mass spectrometry (MS), nuclear magnetic resonance (NMR) or infrared (IR) spectra of the unknown. The second problem is much more challenging and supposes that the unknown is a novel compound synthesized or isolated for the first time. The methods of solving these two problems are quite different. Therefore the first step is the assignment of a given unknown to one of the two mentioned categories. As we will see later in Section 1.6 specific procedures are used for this goal. This book will focus primarily on the structure elucidation of the new organic compounds.
The problem of elucidating the structure of a new compound can be divided into the two following sub-problems: (a) establishing the molecular formula, i.e. determine the type and number of each of the chemical elements making up the molecule, and (b) determining how the atoms are connected by chemical bonds of different multiplicities in the structure. Modern approaches for the determination of molecular formula are well established and based on high-resolution MS (HRMS). In general, they allow for the unambiguous determination of the molecular formula of an unknown.5
Molecular structure determination is such a challenge primarily due to the phenomenon of isomerism. In the 18th century Alexander Humboldt was probably the first who conjectured that there might be chemical substances which are composed of the same set of atoms but have different properties. This was later proven experimentally by Gay-Lussac, Liebig and Wohler6 at the beginning of the 19th century, and the new term “isomers” was introduced into chemistry by Berzelius7 in 1830. The following questions then became of interest to chemists: how many isomers can theoretically exist for a given molecule and how can they be exhaustively enumerated? The mathematical challenge of enumerating all isomers corresponding to a specific molecular formula was later realized when the notion of atom valence was defined and the first research into this area was initiated by Cayley8 in the second half of the 19th century. The computation of the number of isomers from the molecular formula became possible only in the 1960s when computers arrived on the scene and computational chemistry, as a specific area of scientific investigation, was born. Mathematical algorithms and programs were then developed9–11 that provided a possibility not only to calculate the number of isomers corresponding to a given molecular formula but also to generate the structural formulas for all isomers, which is clearly an important capability. As a result, chemists then had the ability to estimate the magnitude of the number of isomers that could be related to well-known substances and the values were rather unexpected. For example, it turned out that benzene was one of 217 conceivable isomers with the composition of C6H6 and articles were published in which all of the structural isomers were enumerated and depicted for the first time.12 The number of isomers produced by the calculations showed that the number is unexpectedly large, even for small molecular formulae.
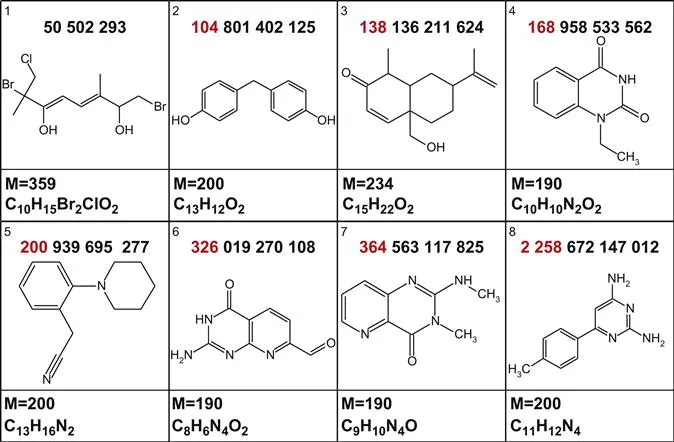
Figure 1.1 displays the structures associated with a series of modest-sized chemical compounds and the number of potential calculated structural isomers.13,14
Figure 1.1 The structures of a series of small molecules and the theoretical numbers of isomers (N) associated with the related molecular formulae.
Figure 1.1 shows that even the simplest of structures can have hundreds of millions, up to trillions, of isomers. For the simple structure with the associated molecular formula of C10H17Br2ClO2, the number of isomers, N, exceeds 50 million and rudimentary inspection suggests that more than 40 million of these could likely exist. It should be noted that the CAS registry15 contains “only” 56 million known chemical compounds while 45 million are commercially available.
The number of isomers associated with the structures of medium-sized complex organic molecules can be estimated as approximately 1020–1030 isomers (in the order of Avogadro's number). At the same time, the following very important conclusion can be drawn: although the number of possible isomers is huge, those corresponding to a given molecular formula do make up a countable(at least in principle) and finite set. With this in mind, we can immediately formulate the following general CASE strategy: to eliminate “superfluous” isomers from the full isomer set by imposing different structural constraints. Figuratively, the general CASE strategy is similar to that of a sculptor who removes superfluous material to produce a masterpiece (Figure 1.2).
Figure 1.2 The analogy between the CASE strategy and the technique of a sculptor.
The more constraints that are imposed, and the more severe they are, then the larger the number of “superfluous” isomers that will be rejected. Since structure identification boils down to the selection of a unique structural formula assigned to an unknown, then a successful result depends on the screening and rejection of N–1 structural formulae that do not comply with the experimental data and constraints applied. We will call the set of n non-identical isomers (1≤n<N) selected as a result of imposing a series of constraints the solution of the structure elucidation problem. The solution is called valid if it contains the correct (genuine) structure and otherwise it is invalid. The solution is called unambiguous if the response file contains only one structure. It should be noted that an unambiguous solution can be either valid or invalid. If the response file contains no structures (n=0), then the imposed constraints are contradictory and the problem has no solution under the chosen conditions. The conceivable constraints leading either to unique structure or at least to a manageable set of plausible structures are outlined below.