Cyclic peptides are increasingly employed as chemical tools in biology and drug discovery. They have gained a lot of interest as alternative sources of new drugs to traditional small molecules.
This book introduces cyclic peptides and provides a thorough overview of biosynthetic and fully synthetic approaches to their preparation. Following an introduction to cyclic peptides, biosynthetic and traditional chemical routes to cyclic peptides are reviewed. Due to their size, their synthesis is not trivial. Recent advances in the incorporation of novel structural units are presented in addition to how synthesis and biological methods can be combined. The chemical analysis of this molecular class is also discussed. Furthermore, chapters detail the progression of cyclic peptides as tools in biology and as potential drugs, providing a future vision of their importance.
In total, this book provides the reader with a comprehensive view of the state-of-the-art of cyclic peptides, from construction to possible clinical utility. This book will be an essential resource for students, researchers and scientists within industry in medicinal, bioorganic, natural product and analytical chemistry fields.

eBook - ePub
Cyclic Peptides
From Bioorganic Synthesis to Applications
- 380 pages
- English
- ePUB (mobile friendly)
- Available on iOS & Android
eBook - ePub
Cyclic Peptides
From Bioorganic Synthesis to Applications
About this book
Trusted by 375,005 students
Access to over 1.5 million titles for a fair monthly price.
Study more efficiently using our study tools.
Information
CHAPTER 1
An Introduction to Cyclic Peptides
Helmholtz-Institute for Pharmaceutical Research Saarland, Department Drug Design & Optimization, Campus E81, Saarbrücken 66123, Germany
*E-mail: [email protected]
1.1 Of Peptides and Proteins (and Small Molecules)
When we compare the Ancient Greek etymologies of ‘protein’ and ‘peptide’ it becomes evident that the former of these very closely related biomolecules is associated with rather positive attributes, as its name derives from proteios (πρωτεῖoς) meaning ″the first quality”.1 Peptides, on the other hand, seem to be considered a mere rudiment of their bigger ancestors, as they are referred to with a terminus derived from peptós (πεπτóς), meaning “digested” or “cooked”.2 But what is the decisive characteristic that defines the ‘quality’ of proteins, which peptides supposedly lack? Both biopolymers usually consist of a linear sequence of amide-linked building blocks, which in most cases are a selection from the standard repertoire of the 20 proteinogenic amino acids. Usually, amino acid sequences shorter than 50 residues are considered as peptides and, thus, these oligomers reside in the so-called ‘middle space’ (see Figure 1.1).3,4
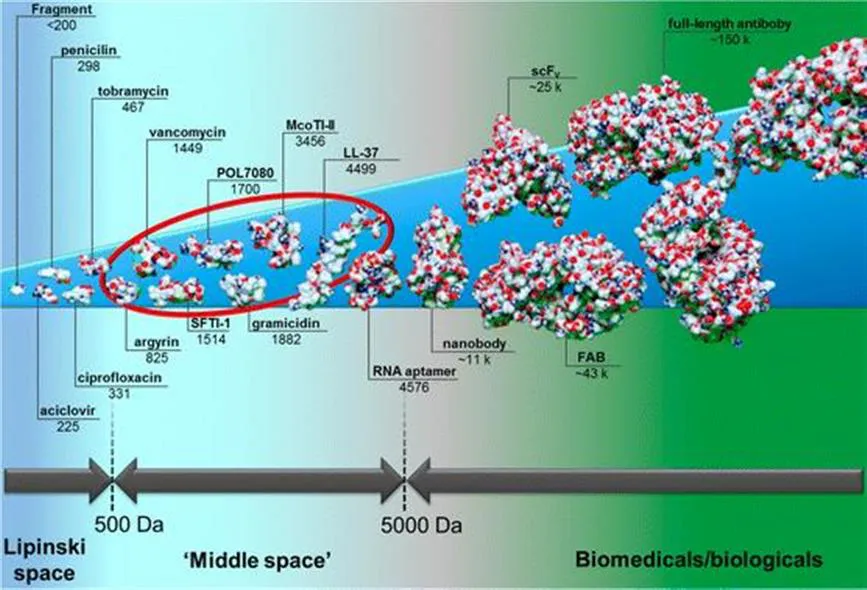
Figure 1.1 The scale of pharmaceutical agents divided into small molecules (Lipinski space), the ‘middle space’, and the biomedicals/biologicals illustrated by selected anti-infective agents and other examples. The range commonly occupied by (cyclic) peptides is highlighted with a red elliptical shape. From small to large: a fragment-sized compound,5 acyclovir, penicillin, ciprofloxacin, tobramycin, argyrin,6 vancomycin,7 sunflower-trypsin inhibitor-1 (SFTI-1),8 POL7080,9 gramicidin,10Momordica cochinchinensis trypsin inhibitor-II (McoTI-II),11 human cathelicidin LL-37,12 an RNA aptamer,13 a vNAR antibody fragment as a representative for ‘nanobodies’,14 a single-chain variable fragment (scFv),15 a FAB-fragment,16 and a full-length IgG antibody.17 The corresponding molecular weights are given in Da.
This term has been coined to refer to molecules with a molecular weight between 500 and 5000 Da (or maybe only approximately 3000 Da). Noteworthily, most of the active principles in pharmaceutical drugs belong either to small molecules or biomedicals.18 The former usually obey the well-known rules-of-five set up by Lipinski, while the most prominent members of the latter are immunoglobulins or derivatives thereof.19 Members of the ‘middle space’ are still comparably rarely found in pharmaceuticals that are in clinical use and small molecular drugs make up about 90% of the pharmaceutical market.20,21 However, their share is continuously growing.20
So, size as a molecular descriptor might give us a hint toward the ‘special quality’ of proteins. However, we know of so-called ‘mini-proteins’ (e.g. McoTI, see Figure 1.1), which can be smaller than 40 residues and exert fascinating biological functions, nonetheless.22,23 Hence, certainly, the line between peptides and proteins is fuzzy and not so clear cut as one might think. Which brings us back to the initial question: ‘What is the unique quality of proteins, then?’
One or maybe THE extraordinary characteristic of protein chains is their ability to adopt precise spatial arrangements of each individual rotatable bond within their backbone. Only when the huge array of so-called Φ, Ψ, and ω dihedral angles is assembled in the right way (see Figure 1.2A), is the protein folded into its correct three-dimensional structure.24 Partial motifs of protein folds (secondary structures) can also be present in peptides (Figure 1.2B).25 The most prominent secondary structures are of course α-helices and β-sheets. Aside from larger loops, which are usually not well defined, β-turns should be highlighted as important structural motifs connecting, for example, the separate strands of a β-sheet.26 However, full tertiary or even quaternary assemblies are usually not found for peptides (Figure 1.2C and D). The reason for this is that a short peptide chain offers fewer opportunities for structure-defining intramolecular interactions, which are needed to render a desired three-dimensional structure thermodynamically favorable. Instead, linear peptides are generally quite flexible and do not adopt an unambiguous geometry.27 As the structure dictates the function of a biomolecule, we may now have approximated the answer to the question raised above: linear peptides often lack a well-defined structure, while proteins exert their various activities through folding into stable three-dimensional assemblies.28
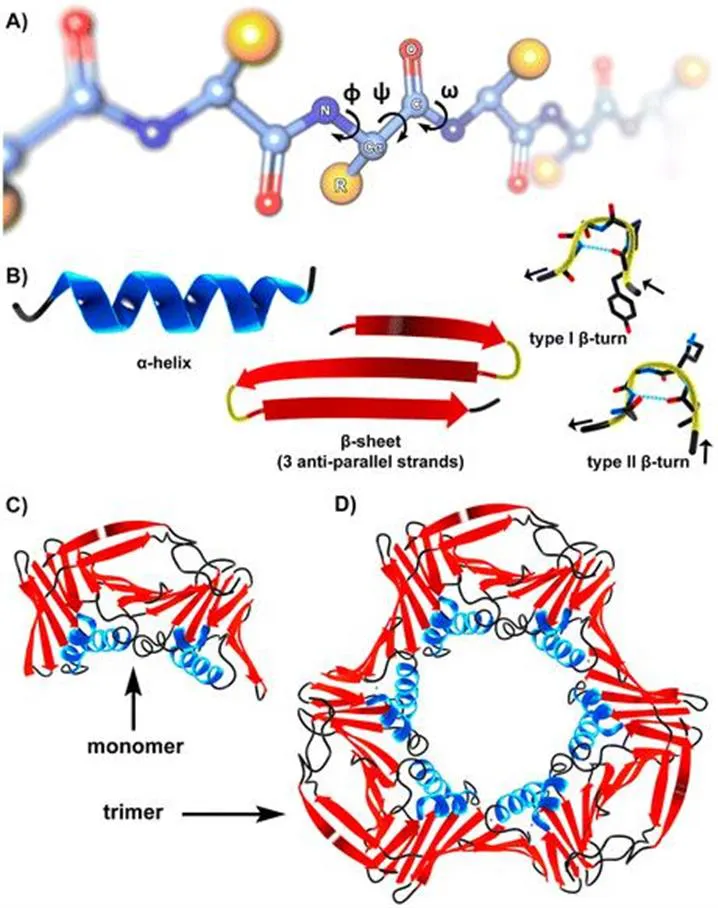
Figure 1.2 (A) A generic peptide chain illustrating the Φ, Ψ, and ω dihedral angles at one amino acid residue. The consecutive order of amino acids in the polypeptide chain is called the primary structure. (B) Schematic depictions of the most common peptide secondary structures: α-helix, β-sheet (3 strands), and type I as well as type II β-turns. (C) and (D) Tertiary and quaternary structure illustrated for the example of proliferating-cell-nuclear-antigen (PCNA).31
This short repetition of textbook knowledge above instantly raises another question: ‘Is there a way to fix a well-defined structure within a peptidic biomolecule?’ Well, the title of the book you hold in your hands readily reveals the answer: Yes! – and it can be achieved through introducing cyclic or macrocyclic motifs! Nature and scientists alike make use of these conformational constraints, which drastically reduce the degree of rotational freedom within the backbone, thereby orienting the side chains in favorable directions and, thus, tethering fascinating biological activities and other favorable properties into otherwise inactive peptide sequences.29,30
1.2 Conformational Constraints
Restricting the conformational freedom of a peptide can be achieved by different means.29 In addition to the just-mentioned macrocyclization strategy, available torsional angles can be reduced through bulky amino acid side chains, where only some of the possible dihedral combinations are allowed at the corresponding and neighboring residues due to steric hindrance.32 Furthermore, the presence of a proline readily fixes the Φ dihedral at this residue as a consequence of the covalent linkage of the alkyl side chain to the α-amine. The ω dihedral, which is commonly referred to as the peptide bond, can be cemented into each of the two possible conformations through biomimetic exchange, for example by the use of 1,2,3-triazoles or other heteropentacycles.33 However, introducing a cycle/macrocycle is the strategy that has the most prominent impact on the overall conformational freedom effecting a multitude of residues and not only some selected positions within the peptide chain.29
There are three straightforward concepts to achieve a looped structure within a peptide:
Head-to-tail-, side chain-to-side chain-, and side chain-to-terminus-cyclization.34,35 Multiple loops are also regularly found in peptides, e.g. a combination of a head-to-tail- and a side chain-to-side chain-macrocycle. If the looped amino acids are only connected via amide bonds, this compound is called ‘homodetic’.35 In cases where any other linkage is involved (e.g. disulfide or depsipeptide bonds), it can be referred to as a ‘heterodetic’ peptide.35 According to the IUPAC, a macrocycle consists of at least twelve atoms. Hence the simplest homodetic macrocyclic peptide is a covalent circuit built up of at least four residues. However, smaller (non-macro-)cyclic motifs also occur containing only a limited number of constrained bonds. Figure 1.3 depicts some of the most simple and common motifs.34 Comparing these structures, it immediately becomes clear that each individual mode of cyclization can result in a completely different geometry of the looped compound.
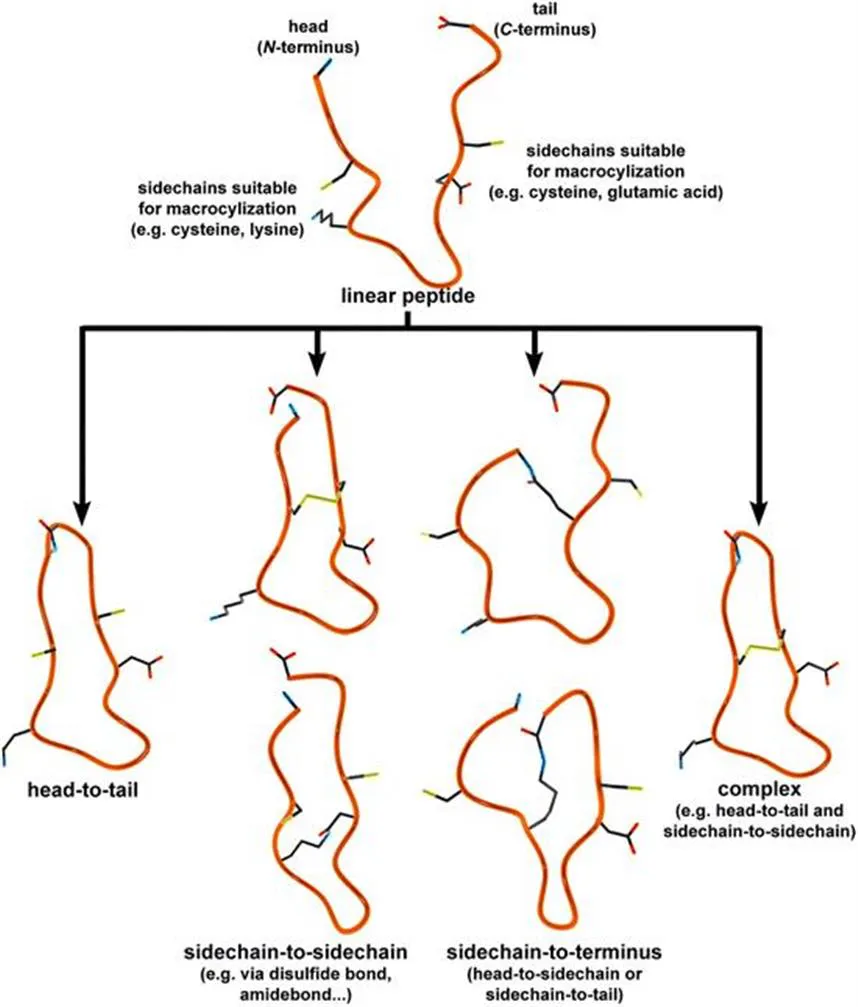
Figure 1.3 Common modes of macrocyclization shown for the example of a generic peptide chain containing two cysteines, a lysine, and a glutamic acid. All other residues are not shown. The peptide backbone is given in orange, carbon in black, nitrogen in blue, oxygen in red, and sulfur in yellow.
Noteworthily, far more complex patterns have been observed in peptidic natural products including, for example, thioether linkages commonly found in lanthipeptides.36 As described for proline, the backbone amide nitrogen can also be used as an anchor point for building a covalent ring...
Table of contents
- Cover
- Halftitle
- Front Matter
- Title
- Copyright
- Contents
- Chapter 1 An Introduction to Cyclic Peptides
- Chapter 2 The Biosynthesis of Cyclic Peptides – RiPPs – An Overview
- Chapter 3 Thioesterase Domain-mediated Macrocyclization of Non-ribosomal Peptides
- Chapter 4 The Biosynthetic Machinery and Its Potential to Deliver Unnatural Cyclic Peptides
- Chapter 5 Modulation of Protein–Protein Interactions Using Cyclic Peptides
- Chapter 6 Biology and Synthesis of the Argyrins
- Chapter 7 Peptide Cross-links Catalyzed by Metalloenzymes in Natural Product Biosynthesis
- Chapter 8 Double-click Stapled Peptides for Inhibiting Protein–Protein Interactions
- Chapter 9 Libraries of Head-to-tail Peptides
- Chapter 10 An Introduction to Bacterial Lasso Peptides
- Chapter 11 Biological Synthesis and Affinity-based Selection of Small Macrocyclic Peptide Ligands
- Chapter 12 Mass Spectrometric Analysis of Cyclic Peptides
- Chapter 13 Experimental and Computational Approaches to the Study of Macrocycle Conformations in Solution
- Chapter 14 Trends in Cyclotide Research
- Chapter 15 Cyclic Peptides – A Look to the Future
- Subject Index
Frequently asked questions
Yes, you can cancel anytime from the Subscription tab in your account settings on the Perlego website. Your subscription will stay active until the end of your current billing period. Learn how to cancel your subscription
No, books cannot be downloaded as external files, such as PDFs, for use outside of Perlego. However, you can download books within the Perlego app for offline reading on mobile or tablet. Learn how to download books offline
Perlego offers two plans: Essential and Complete
- Essential is ideal for learners and professionals who enjoy exploring a wide range of subjects. Access the Essential Library with 800,000+ trusted titles and best-sellers across business, personal growth, and the humanities. Includes unlimited reading time and Standard Read Aloud voice.
- Complete: Perfect for advanced learners and researchers needing full, unrestricted access. Unlock 1.5M+ books across hundreds of subjects, including academic and specialized titles. The Complete Plan also includes advanced features like Premium Read Aloud and Research Assistant.
We are an online textbook subscription service, where you can get access to an entire online library for less than the price of a single book per month. With over 1.5 million books across 990+ topics, we’ve got you covered! Learn about our mission
Look out for the read-aloud symbol on your next book to see if you can listen to it. The read-aloud tool reads text aloud for you, highlighting the text as it is being read. You can pause it, speed it up and slow it down. Learn more about Read Aloud
Yes! You can use the Perlego app on both iOS and Android devices to read anytime, anywhere — even offline. Perfect for commutes or when you’re on the go.
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Please note we cannot support devices running on iOS 13 and Android 7 or earlier. Learn more about using the app
Yes, you can access Cyclic Peptides by Jesko Koehnke, James Naismith, Wilfred A van der Donk in PDF and/or ePUB format, as well as other popular books in Sciences biologiques & Biochimie. We have over 1.5 million books available in our catalogue for you to explore.