![]()
SECTION 1
Human Arylamine N-Acetyltransferases
![]()
Chapter 1.1
Drug Metabolism and Pharmacogenetics Then and Now
Edith Sim*,† and Nicola Laurieri*,‡
Abstract
Arylamine N-acetyltransferases (NAT, EC 2.3.1.5) were amongst the first enzymes known to result in a genetically determined response to a drug, in this case isoniazid. We now know there are two functional NAT genes in humans and these encode (HUMAN)NAT1 and (HUMAN)NAT2 enzymes. The NAT genes are each encoded at polymorphic loci. This chapter explores the early studies on phenotypic variation in NAT activity and discusses how genotyping of (HUMAN)NAT1 and (HUMAN)NAT2 has changed since NAT pharmacogenetics were first described 50 years ago. The relationship between clinical and molecular studies on genes and proteins has provided current understanding of the NAT enzymes and their genes in humans, where evidence indicates (HUMAN)NAT1 has an endogenous role in addition to a drug metabolising role. NATs in other animal species, bacteria and fungi, are also introduced and polymorphisms in other species are described. Enzymic and later structural studies established that the acetylation reaction involved the transfer of an acetyl group to a cysteine sulphydryl group, activated as part of a juxtaposed catalytic traid with aspartate and histidine in all NAT enzymes with only one exception in bacteria to date. The body of research on NATs has laid the foundation for an exciting period to come encompassing whole genome analyses and understanding epigenetic control to allow exploitation of NAT biology in order to understand and perhaps treat disease.
Keywords
Toxicity, Drug Metabolism, Tuberculosis, Isoniazid, Acetylation, Catalytic Triad, Acetyl CoA, Arylamine, Gene Polymorphism.
Introduction
The well-established field of pharmacogenetics is a direct precursor of personalised or stratified medicine (1, 2). Many of the ideas in pharmacogenetics rely on advances in understanding drug metabolism, including N-acetylation. Arylamine N-acetyltransferases (NATs) are amongst the earliest examples of pharmacogenetic variation described over 50 years ago (Fig. 1). These advances were made studying treatment of tuberculosis and, fittingly, understanding NAT at the genomic and molecular level has led to new avenues for treating tuberculosis, through the identification of an active NAT in mycobacteria which is essential for survival of the organism inside host macrophages. This latter aspect is covered in the Chapters of Section 3 and Chapter 4.3 of this volume.
This chapter summarises development of our understanding of genetic variation in NATs in humans and introduces many of the themes in this monograph including animal models (see Chapters of Section 2 and also Chapter 1.5), which have influenced understanding of the pattern of genes and pseudogenes encoding NATs and their variant alleles in humans. The relationship between NATs and susceptibility to toxicity, cancer and other conditions are also themes of individual chapters (see Chapters of Sections 1 and 4). Readers are also directed to a review which will expand on the topics introduced here (5). The nomenclature of the NAT enzymes, their genes and variants are discussed in the Epilogue.
Identification of Pharmacogenetic Variation in NAT
Identification of two functional NAT isoenzymes was the result of an accumulation of studies and the key players are shown in Figure 2. There were several synergistic strands of research: in carcinogenesis and cancer susceptibility, toxicology and pharmacology, molecular enzymology and also interest in understanding the relationship between N-acetylation of serotonin (an alkylarylamine) and arylamine N-acetylation. Investigating both O- and N-acetylation was a key question in carcinogenesis which led to real understanding at the molecular level (Fig. 3A). An outstanding review (6) highlighted apparent discrepancies in animal models of fast and slow acetylation compared to data from clinical pharmacology. Controversies arose as the substrate specificities and the number of different isoenzymes in a range of animals were not fully understood and are continuing to be refined (7). It should also be noted that there are mammals which do not have nat genes and these are the canids (8) and also suncus (an Asian mole) (9).
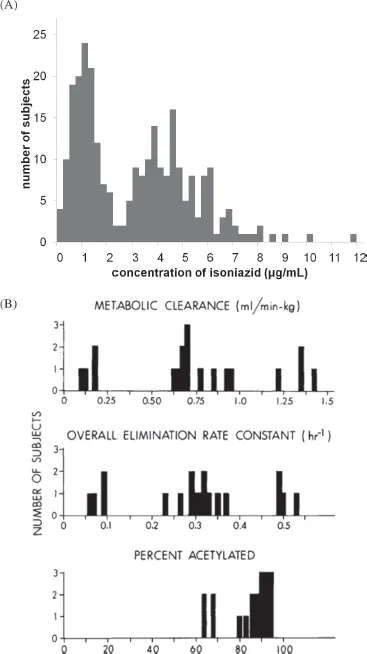
Figure 1: Variation of isoniazid inactivation amongst individuals. (A) Preliminary study showing evolution of plasma isoniazid concentrations 6 hours after drug ingestion amongst 267 patients. All these subjects received approximately 9.8 mg of isoniazid per kg of body weight. Data and figure modified from (3). (B) Distribution of various kinetic parameters of sulfamethazine in 19 subjects. Copyright © 1980 John Wiley and Sons, Inc. Reproduced with permission from (4).
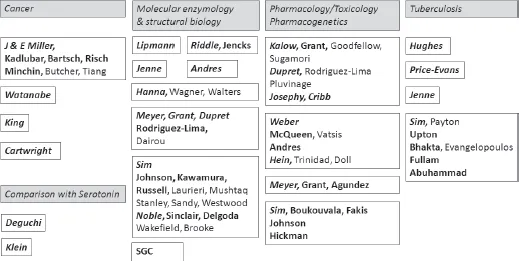
Figure 2: A selection of the main avenues of research in NAT. Group leaders are indicated in bold italics, whilst group members who went on to become group leaders are indicated in bold. Each box contains the names of key researchers in a group. The Millers worked at the McArdle Laboratory and were active from the 1940s. The work on tuberculosis started in the 1950s as Kalow did in his early pharmacogenetics studies. Other teams are described in the text. The vertical order in each subject category indicates the approximate chronological order of publications. SGC: Structural Genomic Consortium.
In the 1950s when tuberculosis treatment was a hot topic and isoniazid was a new drug, studies in monkeys identified acetylisoniazid in urine as the main metabolite (10), in the same year as Lipmann was awarded the Nobel prize in medicine (11, 12) for the identification of acetyl coenzyme A (CoA) as the cofactor in acetylation reactions including arylamine N-acetylation. Interindividual variation in isoniazid metabolism and the terms rapid, intermediate and slow had begun to be used. The possibility of autosomal recessive inheritance of slow acetylation was raised (13) which was subsequently confirmed in a family study (3). The anti-bacterial sulfamethazine, a sulphonamide which is also N-acetylated, was found to follow the same pattern of clearance as isoniazid. Rapid and slow phenotypes were considered as homogygotes (Fig. 1), whilst the intermediate phenotype was a reflection of heterozygosity (4). It is now clear that there is variation in the level of activity associated with different slow alleles. Interestingly, extremely slow outliers were omitted from the graphs in these early studies (3, 13) but may well reflect specific genotypes. The variation in slow acetylation is likely to be related to the mechanism, whereby slow acetylation arises such as generation of unstable unfolded protein or a modification in the shape and enzymic efficiency of the active enzyme. More recent studies are covered below and the early studies on isoniazid are covered in a comprehensive review (14).
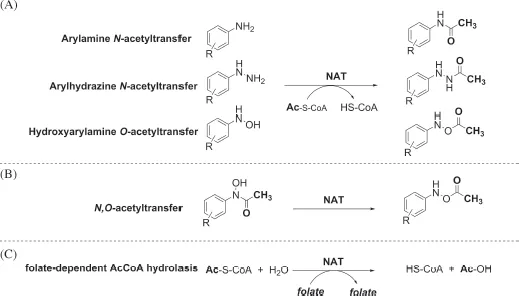
Figure 3: Chemical reactions catalysed by NATs (EC 2.3.1.5). (A) The acetylation reactions involve the transfer of an acetyl group from acetyl Coenzyme A (CoA) to the acceptor. Hydrolysis of acetyl CoA to CoASH also occurs and the rate of hydrolysis of acetyl CoA is greater in the presence of an arylamine acceptor and either acetylation of acceptor or hydrolysis of acetyl donor can be used to measure enzymic activity. Ac: acetyl (B) The N,O-transfer reaction generating an acetoxyester does not involve acetyl CoA. Both O-acetylation and N,O-acetyl transfer are activating reactions leading to carcinogen formation (15). (C) Folate-dependent hydrolysis of acetyl CoA by NATs directly releases acetic acid (AcOH) into the bulk solvent Ac: acetyl. Data from (16).
The inactivation of a different anti-tubercular drug, namely p-aminosalicylate (pAS), was also demonstrated to be due to acetylation. However, the rate of inactivation of these two anti-tubercular drugs, isoniazid and pAS, did not vary amongst individuals in the same way (14). The enzymic activities for N-acetylation of isoniazid and pAS could be separated chromatographically from human liver samples (17) and suggested the involvement of two distinct NAT enzymes; however, it was 25 years later that molecular biology techniques identified that there were two functional NAT genes in humans encoding different enzyme molecules with different specificities (18, 19). We now know that isoniazid and sulfamethazine are substrates for (HUMAN)NAT2 (so-called polymorphic substrates) an...