AN AUTHORITATIVE SURVEY OF CURRENT RESEARCH INTO CLINICALLY USEFUL CONVENTIONAL AND NONCONVENTIONAL ANTIBIOTIC THERAPEUTICS
Pharmaceutically-active antibiotics revolutionized the treatment of infectious diseases, leading to decreased mortality and increased life expectancy. However, recent years have seen an alarming rise in the number and frequency of antibiotic-resistant "Superbugs." The Centers for Disease Control and Prevention (CDC) estimates that over two million antibiotic-resistant infections occur in the United States annually, resulting in approximately 23,000 deaths.
Despite the danger to public health, a minimal number of new antibiotic drugs are currently in development or in clinical trials by major pharmaceutical companies. To prevent reverting back to the pre-antibiotic era—when diseases caused by parasites or infections were virtually untreatable and frequently resulted in death—new and innovative approaches are needed to combat the increasing resistance of pathogenic bacteria to antibiotics.
Bacterial Resistance to Antibiotics – From Molecules to Man examines the current state and future direction of research into developing clinically-useful next-generation novel antibiotics. An internationally-recognized team of experts cover topics including glycopeptide antibiotic resistance, anti-tuberculosis agents, anti-virulence therapies, tetracyclines, the molecular and structural determinants of resistance, and more.
- Presents a multidisciplinary approach for the optimization of novel antibiotics for maximum potency, minimal toxicity, and appropriated degradability
- Highlights critical aspects that may relieve the problematic medical situation of antibiotic resistance
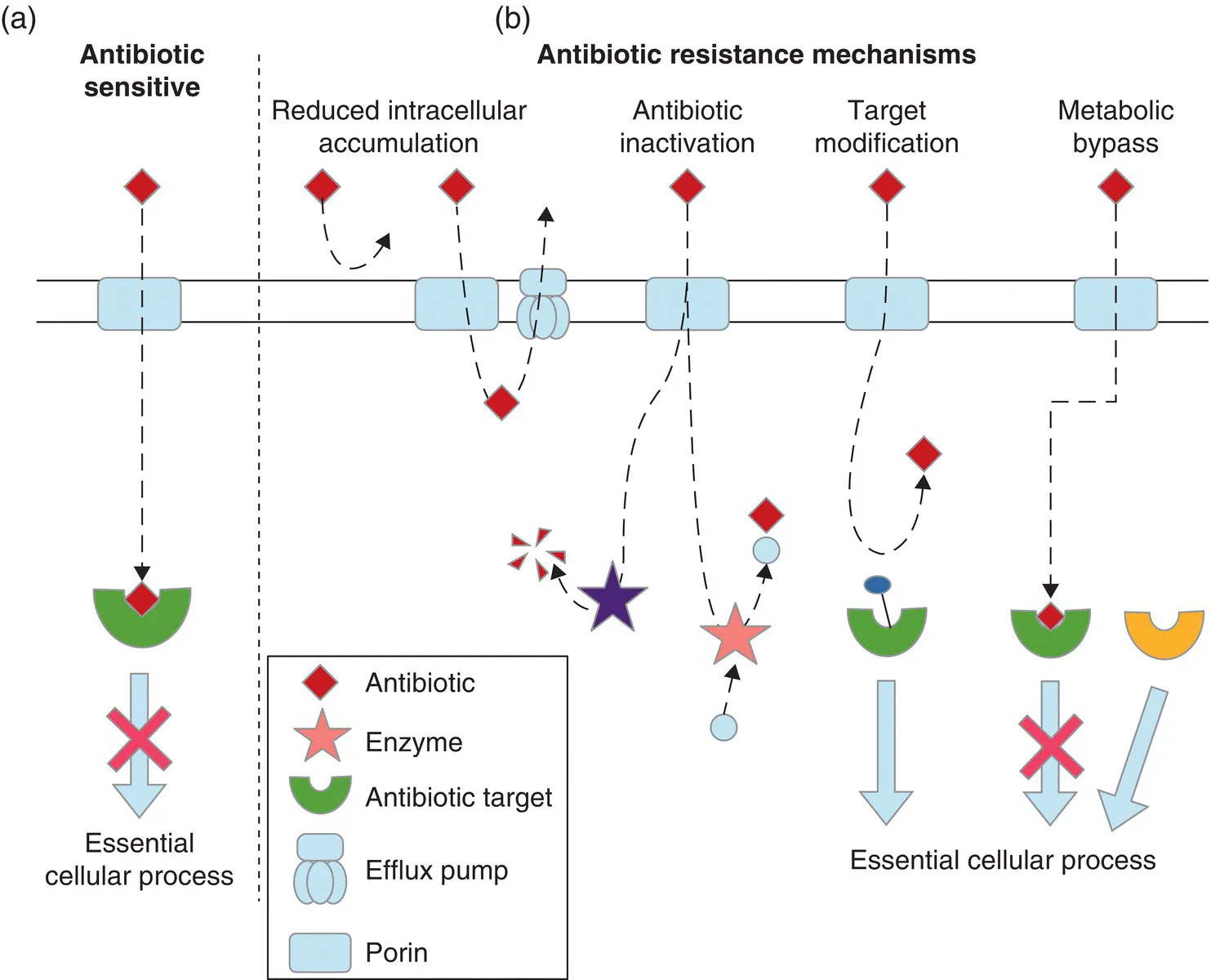
- Includes an overview of the genetic and molecular mechanisms of antibiotic resistance
- Addresses contemporary issues of global public health and longevity
- Includes full references, author remarks, and color illustrations, graphs, and charts
Bacterial Resistance to Antibiotics – From Molecules to Man is a valuable source of up-to-date information for medical practitioners, researchers, academics, and professionals in public health, pharmaceuticals, microbiology, and related fields.